Hi everyone! My name is Wendi Qu and I’m finishing my third year in U of T, majoring in Statistics and Molecular Genetics. I did a ROP399 research project with Dr. Pascal Tyrrell from September 2018 – April 2019 and I would love to share it with you!
Artificial intelligence, or AI, is a rapidly emerging field becoming ever so popular nowadays, with exponentially increasing research published and companies established. Applications of AI in numerous fields has greatly improved efficacy and convenience, including facial recognition, natural language processing, medical diagnosis, fraud detection, just to name a few. In Dr. Tyrrell’s lab in the Department of Medical Imaging, the gears have been gradually switched from statistics to AI in the past two years for research students. With a Life Science and Statistics background, I’ve always been keen on learning the applications of statistics/data science in various medical fields to benefit both doctors and patients. Having done my ROP299 in Toronto General Hospital, I realized how rewarding it was to use real patient data to study disease epidemiology and how my research can help inform and improve future surgical and clinical practices. Therefore, I was extremely excited when I found out Dr. Tyrrell’s lab and really grateful for this amazing opportunity, where I can go one step further and do AI projects in the field of medical imaging.
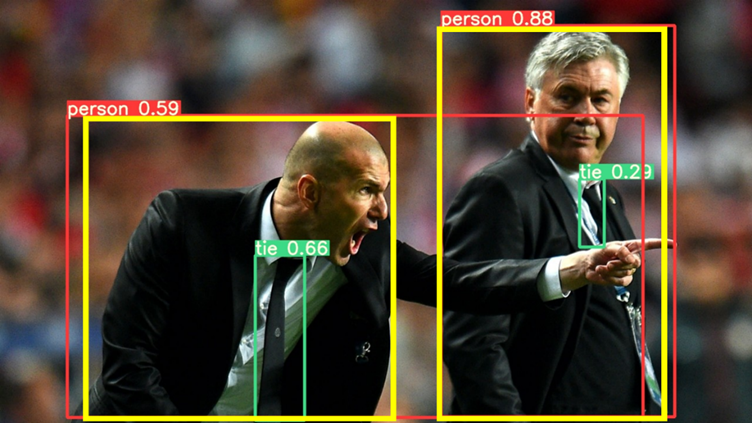
Specifically, my projects focused on how to mitigate the effect of one of the common problems in machine learning – class imbalance. So, what is machine learning? Simply put, we feed lots of data to a computer, which has algorithms that find patterns in those data and use such patterns to perform different tasks. Classification is one of the common machine learning tasks, where the machine categorizes data to different classes (eg. categorizes an image to “cat” when shown a cat image). A common problem in medical imaging and diagnosis is that there’s way more “normal” data than “abnormal” ones. A machine learning model predicts more accurately when trained on more data, and the shortage of “abnormal” data, which are the most important ones, can impair the model’s performance in practice. Hence, finding methods to address this issue is of great importance. My motivation for doing this project largely comes from how my findings can offer insights on how different methods behave when training sets have different conditions, such as the severity of imbalance and sample size, which can be potentially generalized and help better implement machine learning in practice.
However, as with any research project, the journey was rarely smooth and beautiful, especially when I started with almost zero knowledge in machine learning and Python (us undergraduate statisticians only use R…). Starting off by doing a literature search, I realized many methods have been suggested to rectify class imbalance, with two main approaches being re-sampling (i.e. modify the training set) and modifying the cost function of the model. Despite many research done on this topic, I found that such methods were almost never studied systematically to assess their effect on training sets of different natures. The predecessor of this project, Indranil Balki, studied the effect of the class imbalance systematically by varying the class imbalance severity in a training set and see how model performance can be affected. Building on this, I decided to apply different methods to such already established imbalanced datasets and test for model improvement. Because more data lead to better performance, I was also curious if there’s a difference in how much different methods can improve the model in smaller and larger training sets.
One of the hardest parts of the project was making sure I was implementing the methods appropriately, and simply writing the code to do exactly what I want it to do. The latter part sounds simple but becomes really tricky when dealing with images in a machine learning context, and is again, even more challenging if you know nothing about Python… ! After digging into more literature, consulting “machine learning people” in the lab (a big shoutout to Mauro, Ahmed, Ariana, and of course, Dr. Tyrrell), I was able to develop a concrete plan, where I implement oversampling methods via image augmentation only when the imbalanced class has fewer images than other classes, and apply under sampling only when imbalanced class has more images; class weights in the cost function will also be adjusted as another method.
However, implementing them was a huge challenge. I self-learned Python by taking courses in Python, machine learning, image modification, random forest model, and anything that’s relevant to my project on Datacamp, a really useful website offering courses in different coding languages. Through this process and using Indranil’s code as a skeleton, I was finally able to implement all my methods and output the model’s prediction accuracy! It was a long, painful process which involved constant debugging, but it was never more rewarding to see the code finally run smoothly and beautifully!
This wonderful journey has taught me many things – not only have I taken my first step in machine learning, it again reminded me of the most valuable part of doing research, which combines independence, creativity, self-drive, and collaboration. Deciding on a topic, finding a gap, developing your own creative solutions, being motivated to learn new things and conquer challenges, and collaborating with intelligent people surrounding you, are the most invaluable experiences for me this year. Finally, I would love to thank all the amazing people in the lab, especially Mauro, whose machine learning knowledge, coding skills and humour were always there with me, and Dr. Pascal Tyrrell, with more questions back to us when we come with a question, enlightening advice, and a great personality. I appreciate
his amazing experience, and it has inspired me to delve deeper into machine learning and healthcare!
Wendi Qu